Prostatype Genomic ClassifierTM Science & Technology Designed to Change the Paradigm
Prostatype Genomics has made significant advancements in the field through the discovery of new biomarkers that offer an unprecedented understanding of a patient’s prostate cancer biology. By evaluating the RNA expression levels of three embryonic stem cell gene predictors – IGFBP3, F3, and VGLL3 – Prostatype® provides invaluable information that cannot be obtained from traditional clinical factors alone. Prostatype combines this proprietary gene expression signature with the patient’s clinical parameters like PSA, Gleason, and Tumor stage to generate a personalized risk analysis. This test is transformative for urologists, providing them with an individualized prognosis of the patient’s tumor aggressiveness, 10-year metastatic risk, and 10-year prostate-cancer-specific mortality risk. This information is crucial in determining a patient’s eligibility for active surveillance and ensuring that they receive the right treatment at the right time.
The Prostatype Genomic Classifier Assesses the Gene Expression Profile of Three Prostate Cancer Stem Cells
Prostatype measures the expression of three embryonic stem cell gene predictors correlated with both overall and prostate cancer-specific survival:
- IGFBP3: IGF-pathway, pro-apoptosis, anti-metastasis in PCa
- F3: tissue factor, involved in tumor progression
- VGLL3: involved in cancer proliferation
Prostatype interrogates these genes to better understand the patient’s unique tumor biology, thereby providing insights that existing clinical pathological factors alone cannot. The genomic results are combined with PSA, Gleason and Clinical T- stage, to calculate the patient’s personalized risk.
The Prostatype Genomic Classifier provides individualized insights into the patient’s cancer aggressiveness, helping guide personalized treatment decisions.

Development of The Prostatype Genomic Classifier Cancer Stem Cell Gene Expression Signature
A study was designed to explore the importance of embryonic stem cell (ESC) gene signatures, we identified 641 ESC gene predictors (ESCGPs) using published microarray data sets.
ESCGPs were selected in a stepwise manner and were combined with reported genes. Selected genes were analyzed by multiplex quantitative polymerase chain reaction using prostate fine-needle aspiration samples taken at diagnosis from 189 prostate cancer patients diagnosed between 1986 and 2001.
Of these patients, there was overall and prostate-cancer-specific survival data available for 97.9%, and 77.9% were primarily treated by hormone therapy only.
Univariate and multivariate Cox proportional hazard ratios and Kaplan-Meier plots were used for the survival analysis and a k-nearest neighbor (kNN) algorithm for estimating overall survival.
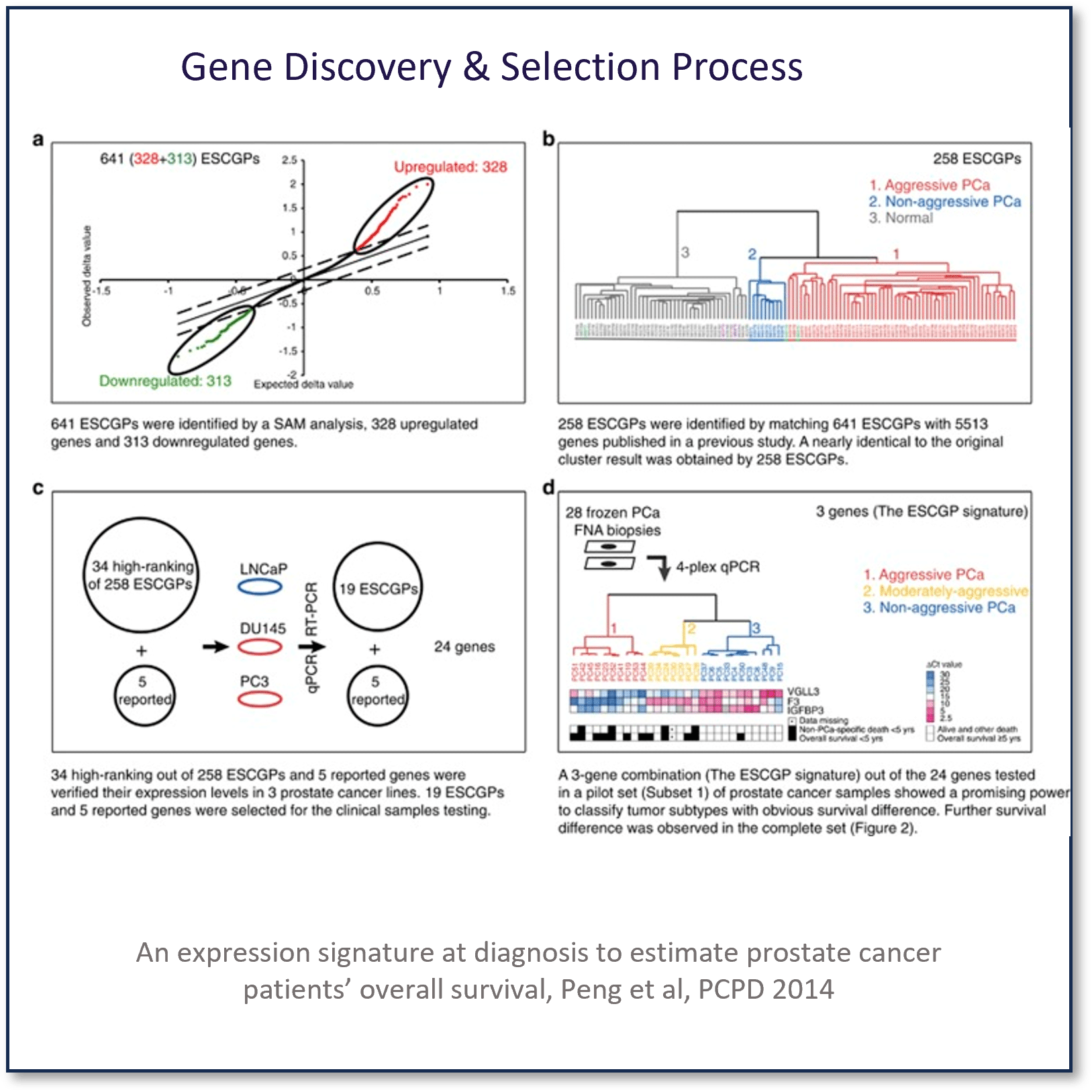
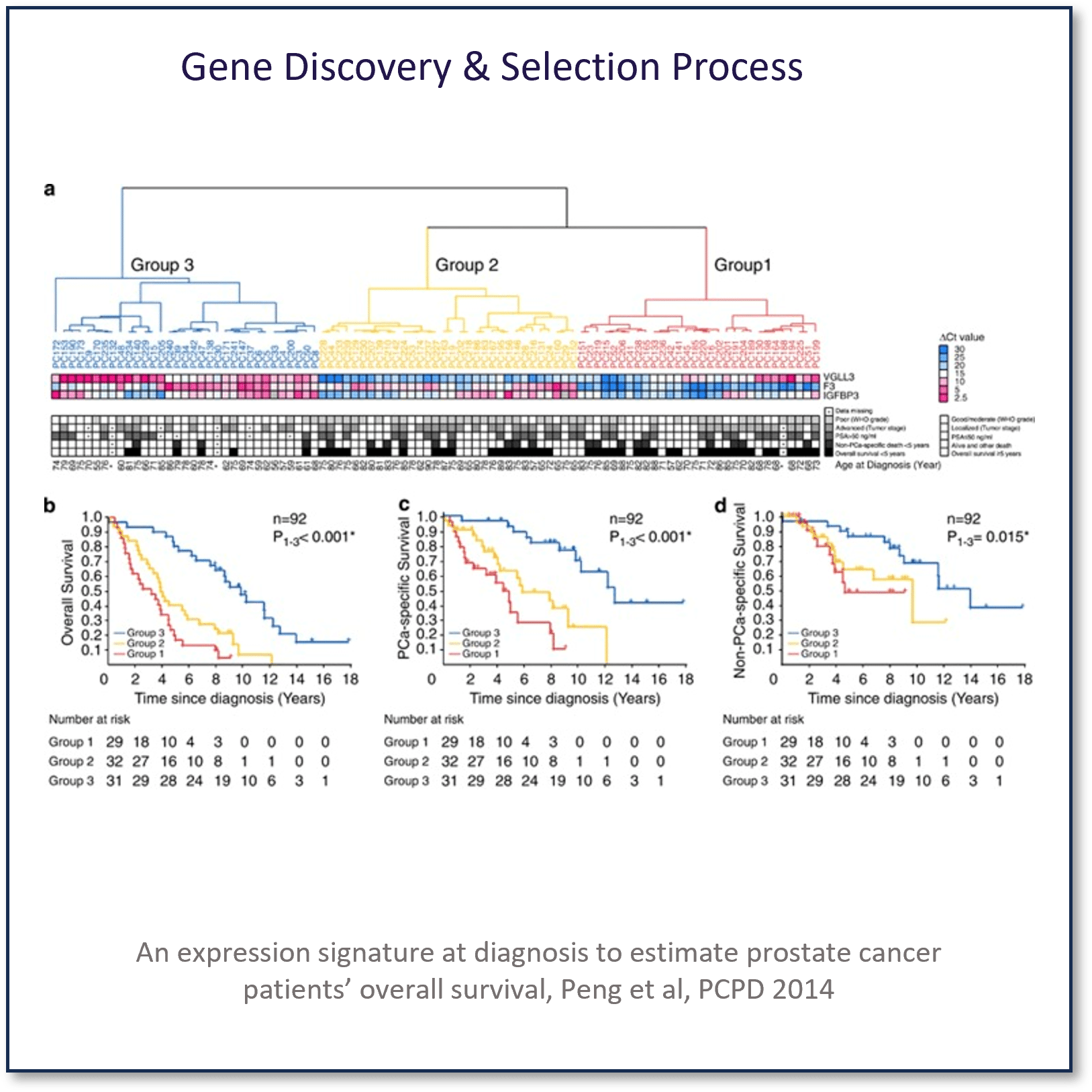
Development of The Prostatype Genomic Classifier Cancer Stem Cell Gene Expression Signature
The median overall survival times of the subtypes were 3.23, 4.00 and 9.85 years, respectively. (This bullet point and the previous do not correlate with regard to risk group and overall survival times.)
The difference corresponded to hazard ratios of 5.86 (95% confidence interval (CI): 2.91-11.78, P<0.001) for the high-risk subtype and 3.45 (95% CI: 1.79-6.66, P<0.001) for the intermediate-risk compared with the low-risk subtype.
The kNN models that included the gene expression signature outperformed the one designed on clinical parameters alone.
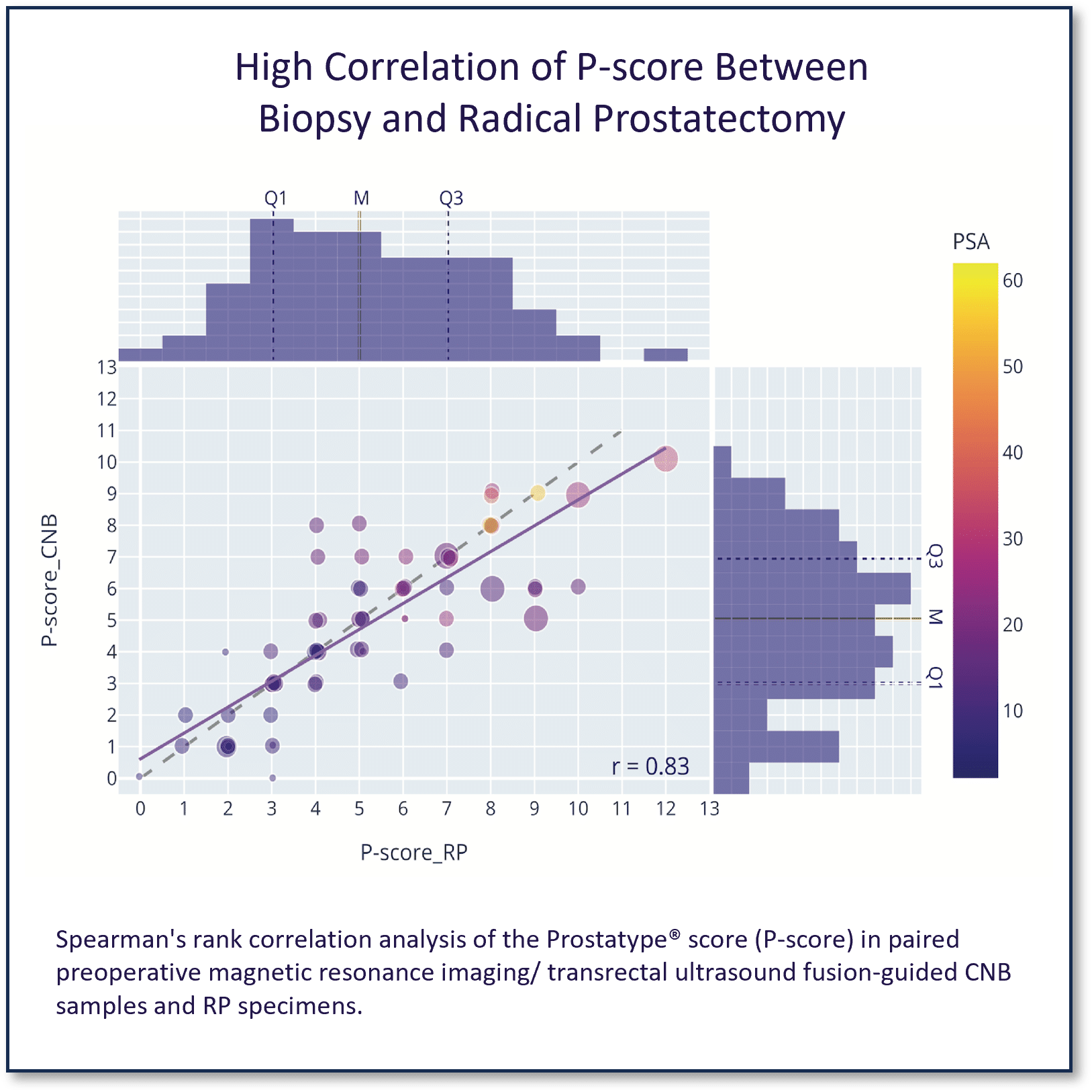
High Correlation of The Prostatype Genomic Classifier between Biopsy and Radical Prostatectomy
This study demonstrated the association between the P‐score measured in preoperative MRI/TRUS fusion‐guided biopsies and the P‐score measured in radical prostatectomy (RP) specimens of PCa patients.
Additional analyses included the association of the P‐score with the pathology of RP specimens and the concordance of the P‐score in paired CNB and RP specimens, as well as in index versus concomitant non-index tumor foci from the same RP was investigated.
The study included 100 patients with localized PCa. All patients were diagnosed by CNB and underwent RP between 2015 and 2018.
Gene expression was assessed with the Prostatype real‐time quantitative polymerase chain reaction kit and the P‐score was calculated. Patients were categorized into three P‐score risk groups according to previously defined cutoffs.
P-Score in preoperative biopsies accurately predicts P-Score in final pathology at radical prostatectomy in patients with localized prostate cancer, Röbeck et al, The Prostate 2023
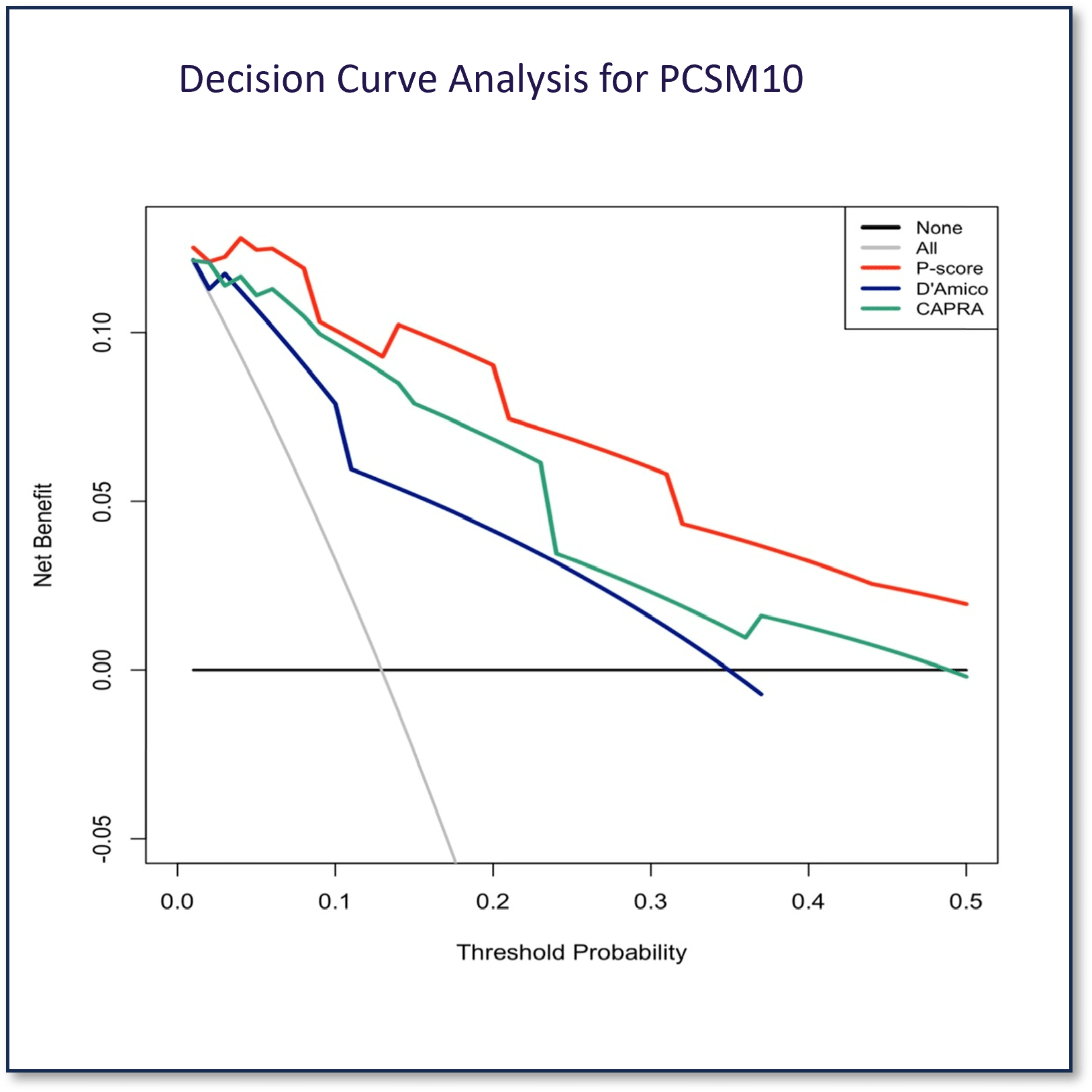
Prostatype Genomic Classifier Decision Curve Analysis Established Prediction of 10-Year PCSM
Sæmundsson et al reported on Prostatype prediction of both 10-year risk for PCSM (HR=1.6) and metastasis (HR=1.46).
Prostatype had an area under curve (AUC) of 0.93 when predicting the PCSM risk at 10 years (95% CI: 0.89-0.98), which was significantly better than both D´Amico (AUC: 0.81, 95% CI:0.72-0.90, p<0.001) and UCSF-CAPRA (AUC: 0.88, 95% CI:0.80-0.96, p<0.05).
Decision curve analysis showed a higher net benefit of the P-score compared to both D’Amico and CAPRA.
For patients who underwent radical prostatectomy (RP), a higher P-score correlated with adverse pathological features such as pathologic tumor stage T3-4 (p<0.0001) and ≥ ISUP 3 (p<0.0001).
Source: Validation of the prognostic value of a three-gene signature and clinical parameters-based risk score in prostate cancer patients. Sæmundsson et al, The Prostate 2023
Before Deciding Know His Prostatype!